Toward understanding life
based on physical principles at molecular level
Students/Researchers
Toward understanding life
based on physical principles at molecular level
2025-06-30 Textbook publication We have published the textbook "生物物理学" by Shoji Takada. Covering a wide range of topics in biophysics, it is an ideal volume for lectures or self-study by undergraduate and graduate students majoring in life sciences, physics, chemistry, and related fields. You can purchace the book here.
More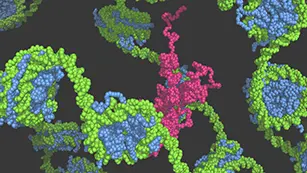
Using multi-scale molecular dynamics simulation techniques, we are working on challenges in the field of molecular cellular biology such as relationship between genome structure and gene expression regulation, operational principles of protein-based molecular motors, molecular dynamics of protein structures. We would like to understand these molecular mechanisms based on physical principles. We developed the coarse-grained molecular dynamics simulator Cafemol.
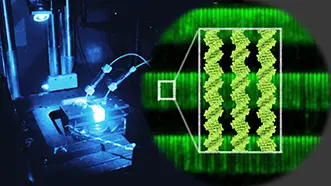
DNA transactions such as replication, transcription, repair, and folding form the basis of life. We are working toward revealing the molecular mechanisms of DNA transactions by combining single-molecule experiments and molecular dynamics simulations.
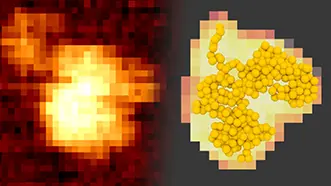
We are developing state-of-the-art information science techniques to enhance space and time resolution of atomic force microscopy images of biomolecules.